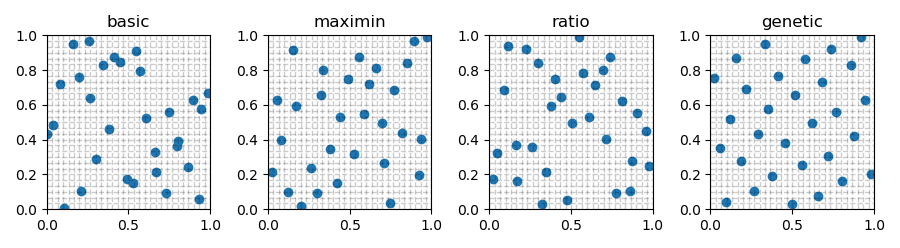
For example, many species distribution models (SDMs), which are used to model how species are distributed along niche gradients, incorporate dispersal kernels to predict range expansions or shifts under different scenarios of environmental change and estimate realistic distributions according to the species’ dispersal potential. the dispersal kernel, is a major determinant of the spatial distribution of individuals, populations and species, thus its accurate estimation is of vital importance for studying and modeling metapopulation and metacommunity dynamics, as well as the distributions and expansion rates of species. The distribution of dispersal distances, i.e.
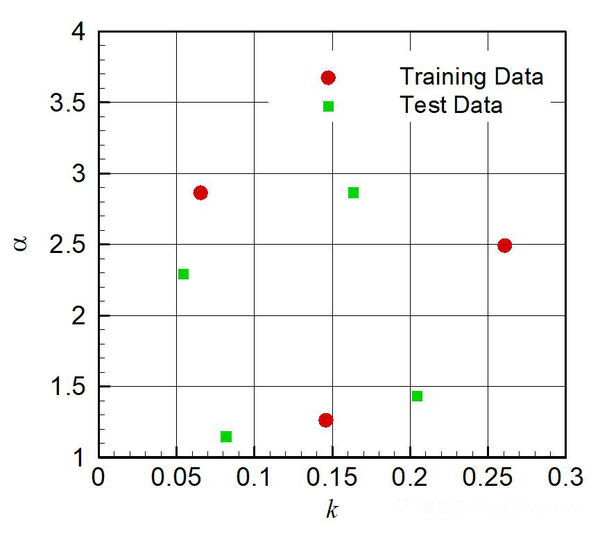
Dispersal distance (D) is usually estimated as the product of the vector movement rate (V) and the retention time (R) of ingested or attached propagules (D = V × R). Propagules are dispersed either externally, entangled in the fur or feathers (“epizoochory” hereafter), or internally, following ingestion and while transiting through the animal’s gut (“endozoochory” hereafter).Ī key element in the study of animal-mediated dispersal is the estimation of the distance at which propagules are dispersed. Among the various vectors, animals such as birds and mammals disperse a great variety of propagules belonging to different species. Instead, they produce dormant propagules that rely on several types of vectors for their dispersal, such as wind, water and animals. Ī considerable part of the Earth’s biota does not actively move. Mechanistic models are widely used in seed dispersal ecology (used here as a general term for the dispersal ecology of dormant propagules, including spores, resting eggs and cysts of plants, animals and fungi), as propagule movement is often difficult to track. The probability distribution of biological variables is of great importance for modeling and understanding biological phenomena, including the mechanistic basis of ecological processes. We also assessed the robustness of these methods to variations in the sampling procedure (sample size and length of sampling time-intervals). To investigate the performance of five different fitting methods, we fitted parametric probability distributions to typical discretized retention-time data with known distribution using as data-points either the lower, mid or upper bounds of sampling intervals, as well as the cumulative distribution of observed values (using either maximum likelihood or non-linear least squares for parameter estimation) then compared the estimated and original distributions to assess the accuracy of each method.
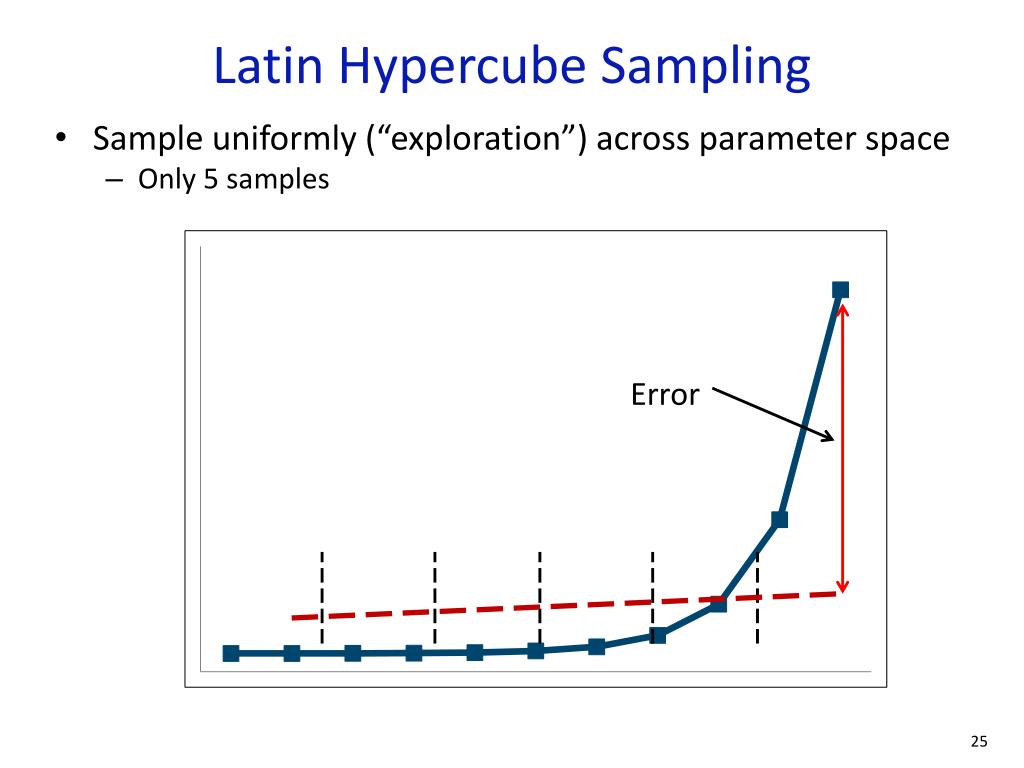
Although parametric continuous distributions have been widely fitted to these interval-censored data, the performance of different fitting methods has not been evaluated. Retention time is a continuous variable, but it is commonly measured at discrete time points, according to pre-established sampling time-intervals. Propagules dispersed by animal vectors are either ingested and retained in the gut until defecation or attached externally to the body until detachment. Propagule retention time is a key factor in determining propagule dispersal distance and the shape of “seed shadows”.
